In part three of our blog post series on the Dimensions API, we’ll have a look at how the Dimensions APIs can help to clean-up, enrich and keep content of a CRIS/RIM system up to date. In case you have missed it, we just recently held a webinar on this topic, that you can find here.
In this blog post we will focus on how the Dimensions APIs can be used to help you with daily tasks around maintaining records and data-cleansing to make your life a little easier:
We’ll take a look at the following three use cases where the enrichment functionality of the Dimensions APIs can assist you:
- Disambiguate: Resolve affiliations and external organization references to unique GRID IDs and complete organization records
- Classify: Automatically classify your content with state of the art, well-known classification systems
- Concepts: Extract keywords from free-text, full-text, abstracts or via DOI
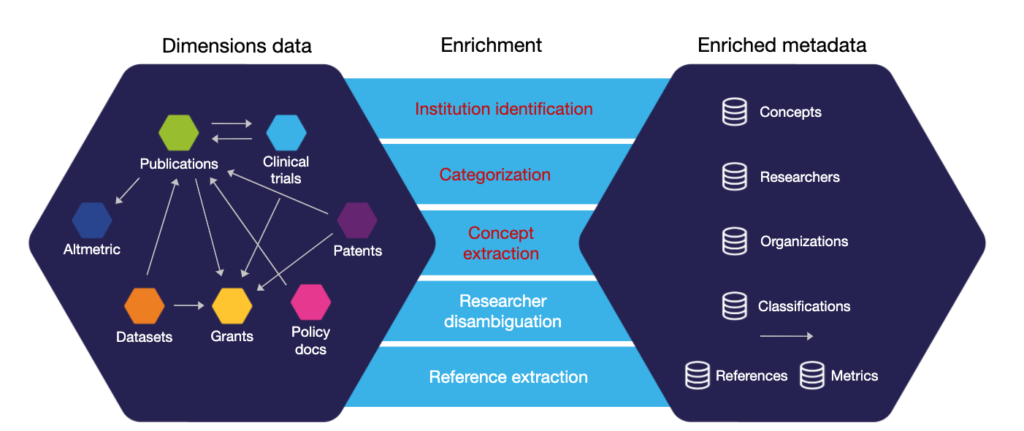
Disambiguate: Untangle affiliation data
We often see that our users have the need to untangle and make sense of various affiliation data. A typical scenario we come across for research information systems is that (unmanaged) organization records usually flow into the CRIS/RIM system when data is imported from various sources or when being keyed in as supporting/supplement data. Usually, the data quality is not high and error-prone resulting in different names, organizations from different hierarchy levels, typos, encoding issues, etc. or simply duplicates. As a result cluttered, unstructured information accumulates over time and becomes a burden; it can even prevent you from accurate reporting, e.g. on collaboration reporting.
To ease the burden of maintaining organization affiliations and making sense of it, the Dimensions APIs can be used to help. With the “extract affiliations” functionality, you can pass structured or unstructured affiliations data/organization information and resolve it to a unique GRID identifier.
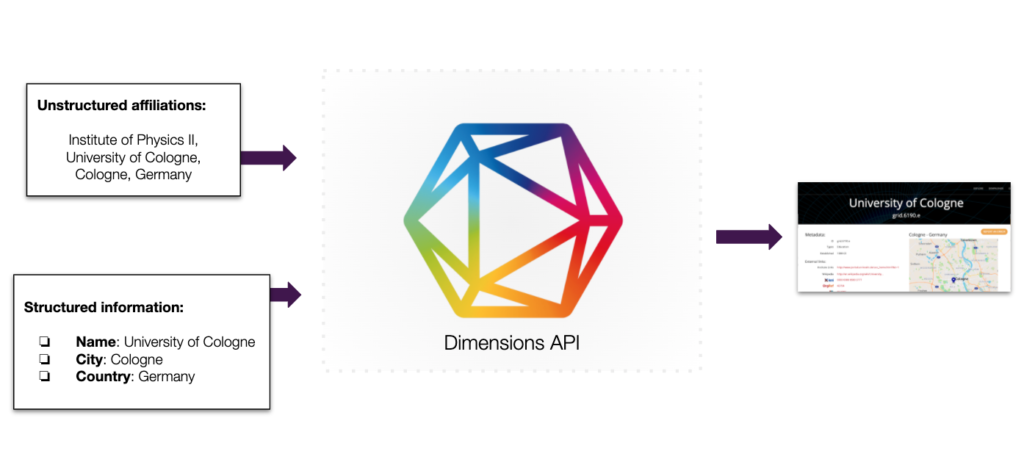
Besides resolving your backlog of existing organization references that are for example derived from co-authorships, collaborations, or awarded grants, records can also be enriched with additional information: like location, organization type, additional IDs, related organizations and much more – all information of the GRID the Global Research Identifier Database is available to enrich your own records.
How well does it work?
We ran a couple of pilots with data derived from CRIS/RIM systems and our results are very promising. We could resolve up to 76% of all source data records, and within these matches, we could identify up to 75% duplicate organizational references. In other words, this feature can be used to resolve a large number of organization records in your systems and potentially uncover a large percentage of duplicate records that currently make your backlog hard to maintain and in most scenarios also more inaccurate.
A possible workflow
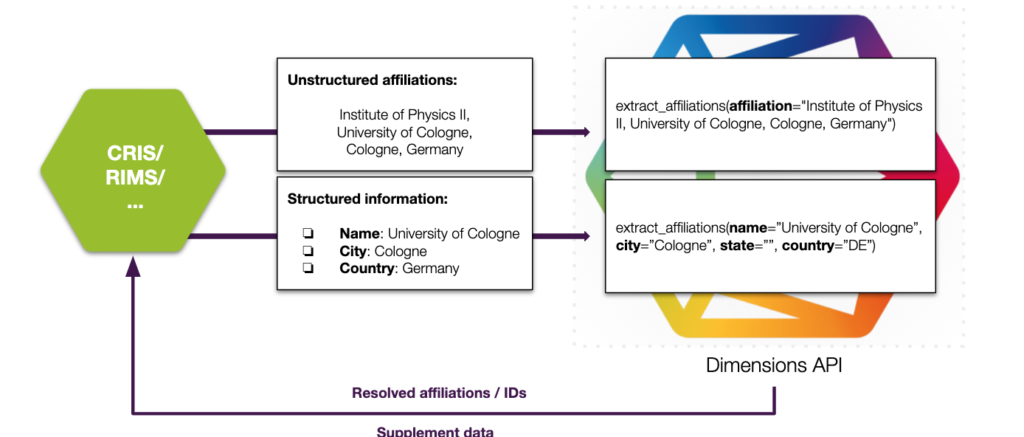
A possible workflow could look like this, where structured and unstructured organization information can be sent to the API and resolved organization records and supplement information to the organization can be retrieved for batch processing or a continuous process.
Classify: Apply classifications to your content
Another enrichment functionality is a feature called classify. It can ingest free text, such as a full-text of a publication, an abstract or any other text string and classify it within a classification scheme of your choosing. The classify functionality works on the fly and can thus be applied in real-time or as a bulk-mechanism to classify all of your existing content. All you need to do is pass your content to the API and choose one of the classification schemes.
An additional benefit is that you don’t necessarily need to have access to the full-text or abstract of a publication. Classifications for content in Dimensions (publications, grants, etc.) can also easily be retrieved by the API via an ID. So in case you don’t have access to full-text but would like to classify your content nonetheless, this can simply be done by provisioning a DOI, a PubMed ID, etc.
A sample workflow could look like this:
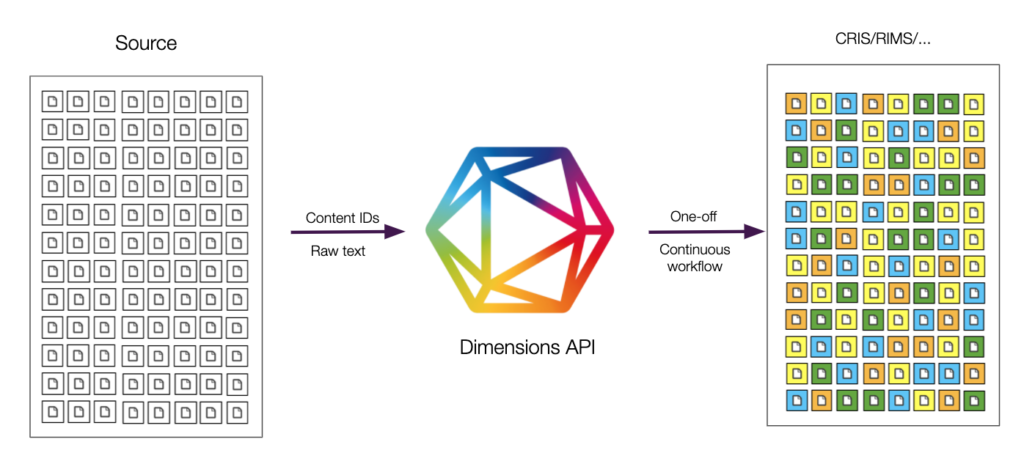
Via an automated approach you can always make sure that content is automatically classified and thus surfaced when searching for a certain field of research.
Dimensions offers a wide-rage of classifications to apply to and enrich your content and new ones are continuously added (like recently the Units of Assessment or the Sustainable Development Goals).
| Area | Classification | From | Granularity |
| All research areas | Fields of Research – FOR codes | Australia/New Zealand, used in national assessment exercise | 176 classification options |
| All research areas | Units of assessment – UoAs | UK, used in REF exercise | 34 classification options |
| Special interest | Sustainable Development Goals (SDGs) | United Nations – developed to put a massive focus on challenges human kind (still) faces | 17 classification options |
| Domain specific | Research, Condition, and Disease Categorization (RCDC) | The NIH uses since 2005 the RCDC classification to report on their funding activities | 295 classification options |
| Domain specific | Health Research Classification System Health Categories (HRCS HC and HRCS RAC) | Two Health related classification with health/disease area and research activity categories, used by UK based funders, development led by MRC | 77 classification options |
| Domain specific | ICRP Common Scientific Outline (ICRP CSO and ICRP CT) | Cancer specific classification developed by ICRP, used by more than 120 funders globally to align | 102 classification options |
You can find the full list of the available research categories and a description here.
Concepts: Extract keywords from any text
Similar to the classify features to categorize content automatically, the concepts functionality uses a modern NLP approach to extract noun phrases. These noun phrases are not tied to entries in thesauri and are domain-independent as a result. Another benefit is the support for emerging terms: the concepts functionality picks up what the author wrote, not what experts agreed should be in the thesaurus. This makes the approach flexible and can be used to automatically extract concepts from any text like a full-text, an abstract of a publication, or simply free-text. In cases where you don’t have access to the full-text, the Dimensions APIs can also be queried with an ID, eg. the DOI of the publication in order to retrieve extracted concepts and keywords:
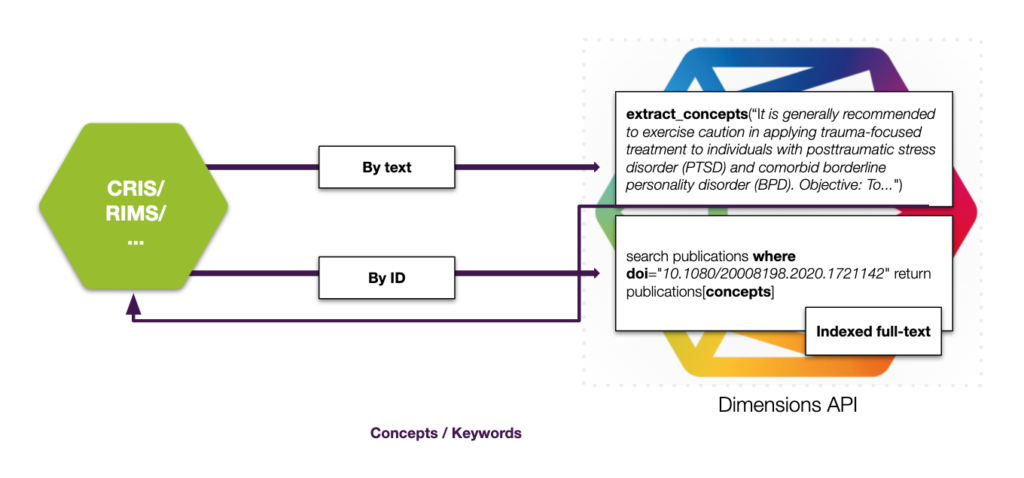
We hope you enjoyed our summary of these three enrichment features of the Dimensions APIs in a CRIS/RIM system scenario, disambiguate, classify and extract keywords. In case you haven’t watched it, we can highly recommend our webinar on the topic with more information and details on the topic.
As always, If you have any questions, please reach out to us
